Inhale, exhale.
Humans breathe between 17,000 to 23,000 times a day, on average. But for more than 70,000 children and young adults around the world, taking a breath can be a struggle because of a rare disease, cystic fibrosis.
A gene that normally triggers a certain protein to move chloride, found in salt, to cell surfaces, where it can attract water, malfunctions in cystic fibrosis patients, so their mucus becomes thick and sticky.
In the lungs, the thick mucus clogs airways. It also becomes a breeding ground for harmful bacteria like Pseudomonas aeruginosa, a particularly nasty pathogen that can quickly lead to inflammation and life-threatening infections.
Normally, mucus is not just a gross byproduct of a cold or viral infection; it is an important biological material that helps us fight off illness. Our bodies make a lot of mucus, maybe as much as 1.5 liters a day, which acts as a protective coating in our sinuses, lungs, intestines, stomach and throat.

“Mucosal layers are such an important barrier between us and the outside world, and we know that any kind of disruption to that is a big source of problems, but really there’s very little known about the interactions between mucins – those primary components of mucus – and bacteria,” said Jason Papin, a professor in the University of Virginia’s Department of Biomedical Engineering who leads a new consortium of engineers and scientists who are using modeling to predict how these interactions might affect a person’s health. Papin’s lab focuses on building computational, predictive models of microbial pathogens.
He is joined by co-principal investigator Roseanne Ford, a professor in UVA’s Department of Chemical Engineering, who has vast experience studying how bacteria move.
Their interdisciplinary, multi-university group recently earned a five-year, $3.2 million grant from the National Institutes of Health to research and build a multi-scale model of mucin interactions, especially with regard to Pseudomonas aeruginosa bacteria. Results of the study could one day lead to better treatments for cystic fibrosis, as well other bacterial infections that involve mucus.
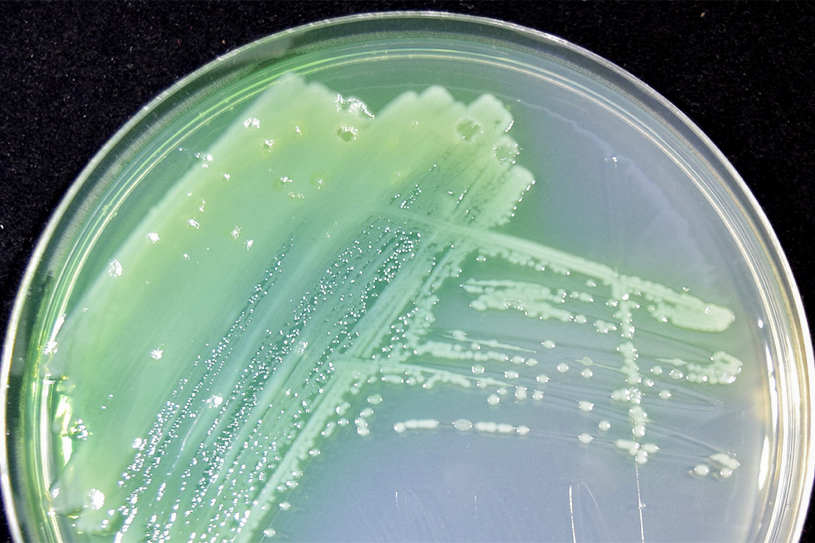
A culture of Pseudomonas aeruginosa, the bacterial pathogen that is the focus of researchers’ NIH grant. This bacterium is the cause of serious health implications in cystic fibrosis patients. (Contributed photo)
In addition to Papin and Ford, the team of researchers includes the UVA School of Medicine’s Shayn Peirce-Cottler, a professor in the Department of Biomedical Engineering specializing in computational and agent-based modeling; Joanna Goldberg, a bacterial geneticist from Emory University, who was formerly a professor in the UVA Department of Microbiology, Immunology and Cancer Biology and has spent decades studying Pseudomonas aeruginosa; and Katharina Ribbeck, Hyman Career Development Career Professor in the Department of Biological Engineering at the Massachusetts Institute of Technology, whose seminal research has focused on mucus barriers.
The new study rose from a pilot grant at UVA Engineering several years ago to develop proposals for large engineering research centers, Papin said. That pilot grant laid the groundwork for the current collaboration and research.
“We came at it from the pathogen, from the microbes, the bacteria,” Papin said. “My lab has done a lot of work with Pseudomonas aeruginosa. This bacterium is a huge problem in the cystic fibrosis community. But it’s a really big problem in lots of other areas like urinary tract infections [and] burn wounds, and people who are immunocompromised, like AIDS patients and cancer patients on chemotherapy.”
Papin has been working with Goldberg for more than 10 years to build computer models of the metabolism of these bacteria, resulting in a greater understanding of how Pseudomonas aeruginosa survives in difficult environments.
“They are these recurrent, long-term lung infections that you can treat with antibiotics,” Goldberg said. “But unlike a typical infection that goes away, this bacterium will continue to come back again and again. And so patients are constantly undergoing treatment with antibiotics.”
One of the things researchers have been studying is how bacteria move in mucus and what kind of chemical signaling might be taking place. Ford has been working with Papin for a couple of years on this and brings her chemical engineering background to bear.
“My role is primarily to look at modeling of chemical transport and bacterial transport at what’s called a macroscopic scale,” Ford said. “Within mucus, the distribution of bacteria in different chemicals affects things like how they might change their phenotype. Depending on those external chemical cues, the cells internally can respond differently by turning on and off different genes, which might make them more or less virulent.”
This grant will enable the researchers to develop a multi-scale computational model that can guide the group’s experimental design with the goal of gaining a better understanding of the relationship between the microbes and the mucin.
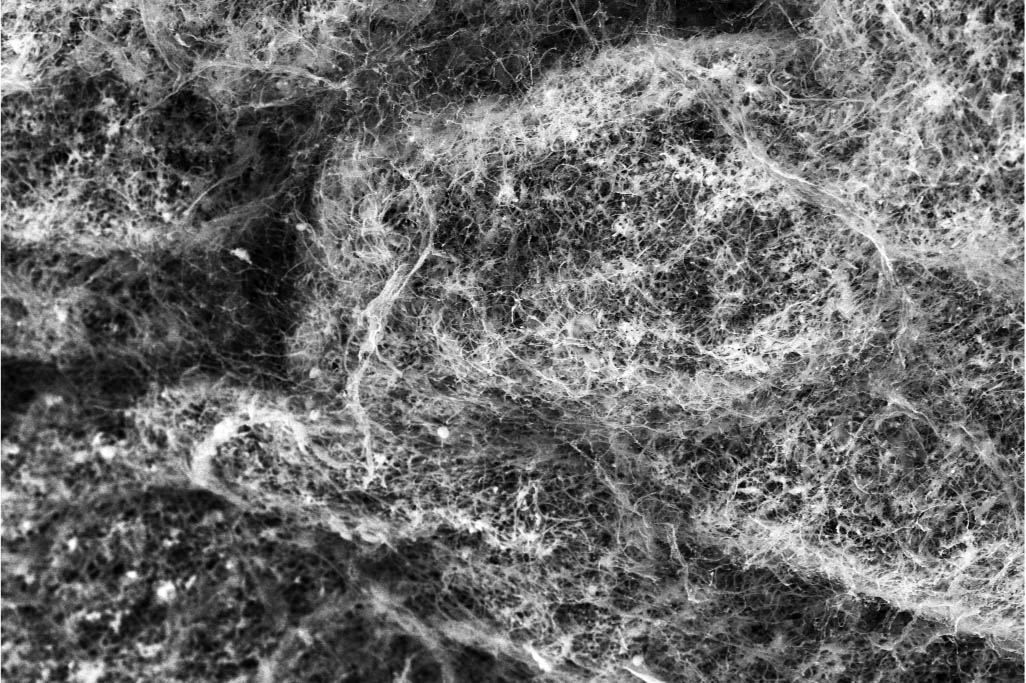

A mucin network, which is the primary component in mucus. (Photo courtesy Katharina Ribbeck)
“Even though we’re focusing on what happens in Pseudomonas aeruginosa, there are millions of other bacteria in the same space, living in the same neighborhood, swimming around in that same mucus,” said Peirce-Cottler, who is co-director of the UVA Center for Advanced Biomanufacturing and an affiliated faculty of UVA’s Fibrosis Initiative. “And so there are interactions between Pseudomonas aeruginosa and other helpful bacteria that live in the mucus to help defend us and combat Pseudomonas aeruginosa. It’s mind-blowingly complicated with respect to these different bugs and these different proteins, but that’s exactly why we must use computational modeling.”
Peirce-Cottler will stitch together the metabolic modeling from Papin and Goldberg with Ford’s chemical transport modeling to produce a predictive model that might allow for more focused treatments for cystic fibrosis and other mucus infections.
The base understanding of mucins and their biological role in our bodies can be traced to Ribbek’s pioneering work; she has been instrumental in defining the biochemistry of mucus.
“Over millions of years, it seems like mucus has evolved the ability to keep problematic pathogens in check. And that was the starting point of my research group, really, to understand how mucus affects the behavior and life of microbes that live inside it,” Ribbeck said.
Her research has revealed a complex ecology that exists within the mucus that can both help us combat pathogens or set up a perfect breeding ground for them. Understanding what all these interactions within mucus mean is challenging.
“You can unpack the complexity of the problem layer by layer with experimental approaches, but then someone needs to put them back together for a comprehensive picture and understanding, and this is what the team aims to accomplish,” Ribbeck said.
Papin believes the research might eventually lead to more effective antibiotic targeting and strategies for treating infections by microbial pathogens such as Pseudomonas aeruginosa.
“We know different people have different susceptibility to different kinds of infections,” Papin said. “So let’s build these computer models to try to capture some of those differences and try to understand what are the weak points in that system so that we can develop new tools and new therapies to improve human health.”
Media Contact
Article Information
July 7, 2020
/content/not-gross-it-sounds-predicting-how-bacteria-mucus-affect-human-health

